Research
During development the cell has access to a toolbox that allows the transformation of transient stimuli into long term changes that define the cellular phenotype. Thus a liver cell “remembers” to be a liver cell. This form of cellular memory is mediated by epigenetic processes such as Histone-modifications, DNA-methylation and non-coding RNAs (Fig. 1).
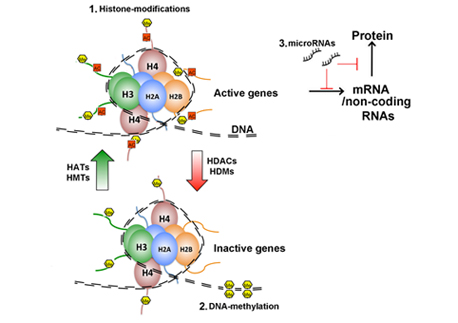
Fig. 1. Epigenetic mechanisms: (1) Dimers of the 4 core histones (H) build the nucleosome around with 147bp of DNA is wrapped. The histone tails are subjected to post-translational modification including acetylation and methylation. Those modifications give rise to a combinatorial pattern, also called the “histone-code,” that affects gene expression. Histone acetylation and methylation is regulated by the counteracting activity of histone acetyltransferases (HATs) and histone deacetylases (HDACs) or histone-methyltransferases (HMTs) and histone-demethylases (HDMs), respectively. (2) Methylation of DNA at the C-5 atom of cytosine is mediated by DNA-methlytransferases and often occurs in Cytosin-Guanin rich regions of the genome (CpG islands). DNA-methylation is generally associated with gene silencing. More recently is has been discovered that Cytosine can be hydroxymethylated which occurs predominantly in brain tissue and compromises a novel mechanism of gene-regulation. (3) Micro RNAs (miRs) are 19-22 nt long non-coding RNAs that catalyze gene-silencing or inhibition of protein translation thereby regulating protein homeostasis.
The same epigenetic machinery contributes to the formation of long-term memories and de-regulation of epigenetic processes is linked with the pathogenesis of complex brain diseases that are driven by variable combinations of genetic and environmental risk factors (GxE interactions). Our lab has pioneered the novel field of neuro-epigenetics. For example, we were the first to apply next-generation-sequencing approaches to the adult brain and could show that the onset of age-associated memory disturbances is mediated by specific changes in histone-acetylation. We could also show that targeting histone-deacetylases or microRNAs ameliorates cognitive decline in models for Alzheimer’s disease. On this basis we proposed a model in which the combination of environmental and genetic risk causes disease-specific epigenetic signatures that drive pathogenesis of complex neuropsychiatric diseases. Identifying such epigenetic signatures would help to understand the mechanisms that underlie memory consolidation and develop individual treatments for brain diseases (Fig.2).
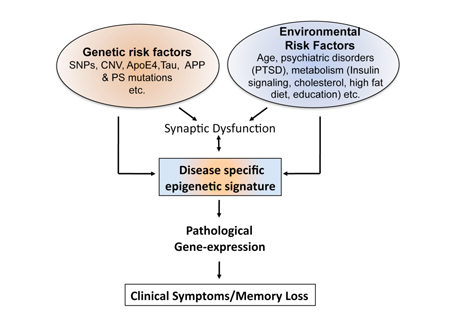
Fig. 2 An epigenetic signature of brain disease. Most data suggest a combination of genetic and environmental factors to be most relevant to the pathogenesis of brian diseases such as sporadic Alzheimer’s disease. Along the course of pathogenesis deregulation of vulnerability genes may occur via multiple mechanisms. Most recent data suggest that epigenetic mechanisms are central to the manifestation of pathological gene-expression and might act as a bottleneck to mediate gene-environment interactions relevant to disease progression. To define disease-specific epigenetic signatures and to identify corresponding therapeutic strategies is a major focus of our research.
Our research program is organized into 4 areas.
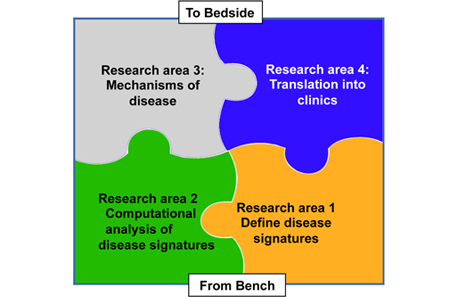
Research area 1: Define disease signatures
We aim to identify disease signatures associated with disease-related phenotypes. Therefore we model environmental and genetic risk factors in suitable animal models including rodents. In a feed-forward feed-backward approach we also analyze disease signatures in well-defined cohorts of human patients. On the basis of this research we will be able to link the onset and progression of diease phenotypes to molecular signatures such as altered epigenome plasticity in specific populations of nerve cells. This will allow us to formulate novel hypothesis on the pathogenesis of sporadic AD and develop novel therapeutic approaches.
Research area 2: Computational analysis of disease signatures
In close collaboration with the Group of Stefan Bonn at the DZNE Göttingen (link to Stefan webpage) we perfomr computational analysis of disease signatures. A specific aim is to understand the dynamics of epigeneitc networks during neurodegenerative diseases and to use machine learning approaches to make predictions of disease progression and therapeutic intervention.
Research area 3: Mechanisms of disease onset and progression
Here we use suitable gain and loss of function approaches to understand the role of disease associated epigenetic modifications and the corresponding enzmyes. These studies form the basis for the design and screening of novel drug candidates.
Research area 4: Translation into clinics, disease marker and therapeutic intervention
In close collaboration with the University Medical Center and the Department for Psychiatry in particular we have established a clinical science platform (Link to Anja’s webpage) that aims to identify and confirm signatures of disease-associated epigenome changes as biomarker and novel drug targets and to perform interventional trials.

